NASA DEM#
NASADEM
This dataset contains an extract of the NASADEM product for the San Diego MSA. NASADEM provides a digital elevation model for the world.
Source: NASA EarthDataURL
Processing: acquisition and transformation of the dataset are documented inbuild_nasadem_sd.ipynb
Build the nasadem_sd extract#
This notebook documents the acquisition and extraction of the San Diego region from the global NASADEM dataset.
%matplotlib inline
import zipfile
import cenpy
import contextily as cx
import rasterio
from rasterio.plot import show as rioshow
import matplotlib.pyplot as plt
Data acquisition#
We use the NASADEM_HGT variant of the project. We select the areas to download through the NASA EarthData Search tool. Once selected, the extract requires the following files:
base_url = ("https://e4ftl01.cr.usgs.gov//"\
"MODV6_Dal_G/MEASURES/NASADEM_HGT.001/"\
"2000.02.11/"
)
f_urls = ["NASADEM_HGT_n33w118.zip",
"NASADEM_HGT_n33w117.zip",
"NASADEM_HGT_n32w117.zip",
"NASADEM_HGT_n32w118.zip"
]
for url in f_urls:
print(base_url + url)
https://e4ftl01.cr.usgs.gov//MODV6_Dal_G/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n33w118.zip
https://e4ftl01.cr.usgs.gov//MODV6_Dal_G/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n33w117.zip
https://e4ftl01.cr.usgs.gov//MODV6_Dal_G/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n32w117.zip
https://e4ftl01.cr.usgs.gov//MODV6_Dal_G/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n32w118.zip
Download is free but requires a login from the USGS. We assume the four files have been downloaded and placed in the same folder as this notebook.
Next is extracting the DEM files (.hgt) and merging them into a single .tif for convenience. .tif is a common file format for rasters that will make it handier later on to work with the merged file to clip it.
Extraction of
.hgtfiles
for zipped in f_urls:
zip_f = zipfile.ZipFile(zipped, "r")
f = zipped.strip("NASADEM_HGT_").strip(".zip")
zip_f.extract(f+".hgt")
zip_f.close()
Merging into a single
.tiffile
! rio merge -f GTIFF *.hgt merged.tif
! rm *.hgt
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
Error: Could not open file : file exists and won't be overwritten without use of the `--overwrite` option.
Extraction of SD region#
Region boundaries
Let’s pull out total population (P001001):
%%time
census10 = cenpy.products.Decennial2010()
sd = census10.from_msa("San Diego, CA",
level="county",
variables=["P001001"]
)\
.to_crs(epsg=4326)
/opt/conda/lib/python3.7/site-packages/pyproj/crs/crs.py:55: FutureWarning: '+init=<authority>:<code>' syntax is deprecated. '<authority>:<code>' is the preferred initialization method. When making the change, be mindful of axis order changes: https://pyproj4.github.io/pyproj/stable/gotchas.html#axis-order-changes-in-proj-6
return _prepare_from_string(" ".join(pjargs))
/opt/conda/lib/python3.7/site-packages/pyproj/crs/crs.py:55: FutureWarning: '+init=<authority>:<code>' syntax is deprecated. '<authority>:<code>' is the preferred initialization method. When making the change, be mindful of axis order changes: https://pyproj4.github.io/pyproj/stable/gotchas.html#axis-order-changes-in-proj-6
return _prepare_from_string(" ".join(pjargs))
CPU times: user 2.61 s, sys: 121 ms, total: 2.73 s
Wall time: 54.4 s
r = rasterio.open("merged.tif")
f, ax = plt.subplots(1, figsize=(9, 9))
sd.plot(facecolor="none", edgecolor="#F93822", zorder=1, ax=ax)
ax = rioshow(r, alpha=0.5, zorder=2, ax=ax)
cx.add_basemap(ax, crs=r.crs)
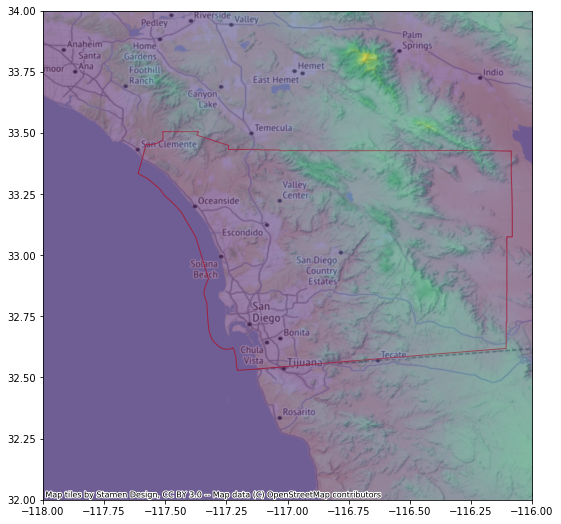
sd.to_file("sd.geojson", driver="GeoJSON")
Clipping of DEM for SD
! rio mask merged.tif \
nasadem_sd.tif \
--crop \
--geojson-mask\
sd.geojson
! rm merged.tif sd.geojson
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
proj_create: Open of /opt/conda/share/proj failed
proj_create: init=epsg:/init=IGNF: syntax not supported in non-PROJ4 emulation mode
Error: Could not open file : file exists and won't be overwritten without use of the `--overwrite` option.
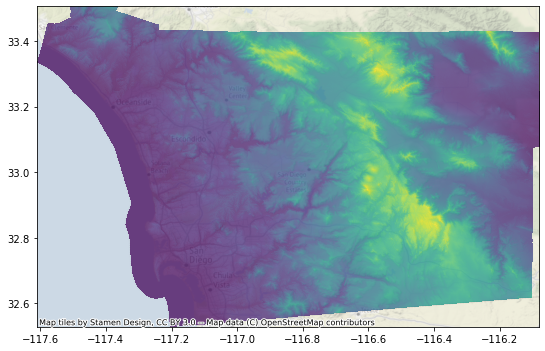
f, ax = plt.subplots(1, figsize=(9, 9))
r = rasterio.open("nasadem_sd.tif")
rioshow(r, zorder=1, alpha=0.75, ax=ax)
cx.add_basemap(ax, alpha=0.5, crs=r.crs)